HLA-A*02:01 presenting "GILEFVFTL" to Alpha/Beta T cell receptor at 2.95Å resolution
Data provenance
Information sections
- Publication
- Peptide details
- Peptide neighbours
- Binding cleft pockets
- Chain sequences
- Downloadable data
- Data license
- Footnotes
Complex type
Class i with peptide and alpha beta tcr
HLA-A*02:01
GILEFVFTL
TRAV27
TRBV19
Species
Locus / Allele group
Molecular basis for universal HLA-A*0201-restricted CD8+ T-cell immunity against influenza viruses.
Memory CD8(+)T lymphocytes (CTLs) specific for antigenic peptides derived from internal viral proteins confer broad protection against distinct strains of influenza A virus (IAV). However, immune efficacy can be undermined by the emergence of escape mutants. To determine how T-cell receptor (TCR) composition relates to IAV epitope variability, we used ex vivo peptide-HLA tetramer enrichment and single-cell multiplex analysis to compare TCRs targeted to the largely conserved HLA-A*0201-M158and the hypervariable HLA-B*3501-NP418antigens. The TCRαβs for HLA-B*3501-NP418 (+)CTLs varied among individuals and across IAV strains, indicating that a range of mutated peptides will prime different NP418-specific CTL sets. Conversely, a dominant public TRAV27/TRBV19(+)TCRαβ was selected in HLA-A*0201(+)donors responding to M158 This public TCR cross-recognized naturally occurring M158variants complexed with HLA-A*0201. Ternary structures showed that induced-fit molecular mimicry underpins TRAV27/TRBV19(+)TCR specificity for the WT and mutant M158peptides, suggesting the possibility of universal CTL immunity in HLA-A*0201(+)individuals. Combined with the high population frequency of HLA-A*0201, these data potentially explain the relative conservation of M158 Moreover, our results suggest that vaccination strategies aimed at generating broad protection should incorporate variant peptides to elicit cross-reactive responses against other specificities, especially those that may be relatively infrequent among IAV-primed memory CTLs.
Structure deposition and release
Data provenance
Publication data retrieved from PDBe REST API8 and PMCe REST API9
Other structures from this publication



Data provenance
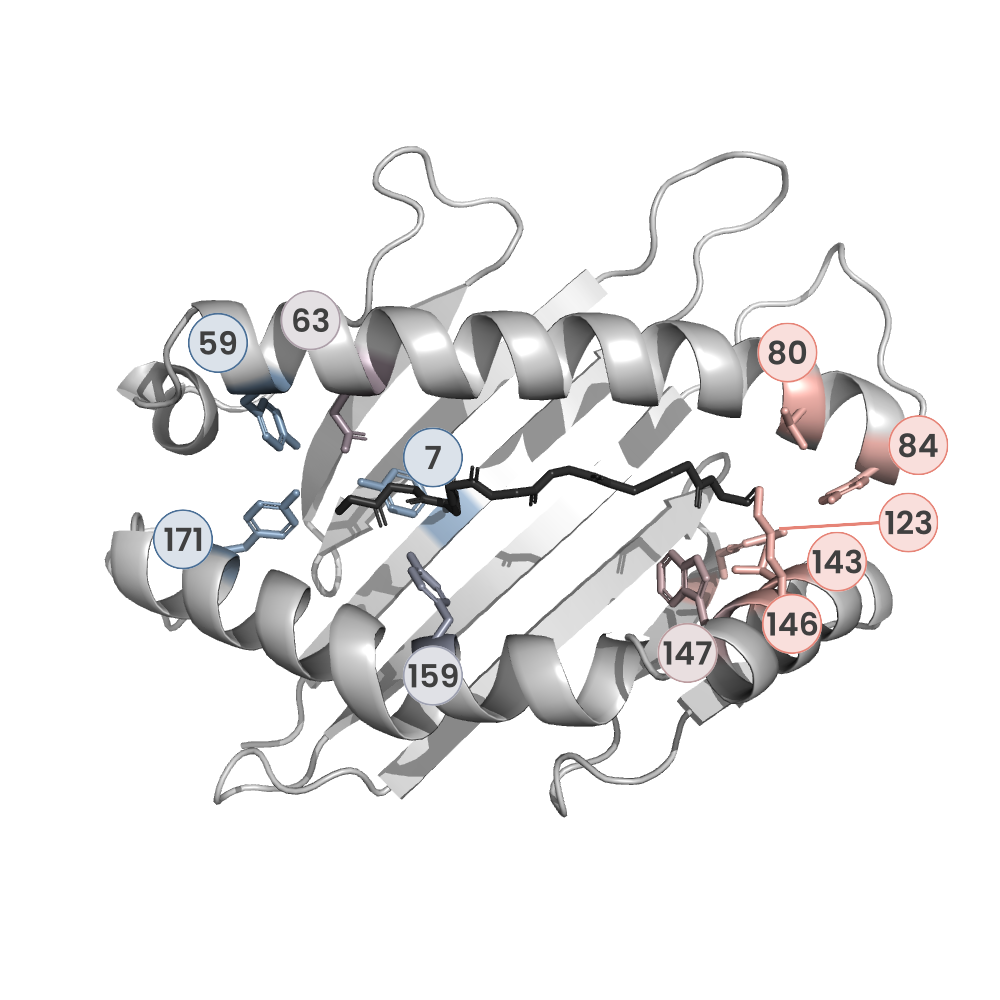
MHC:peptide complexes are visualised using PyMol. The peptide is superimposed on a consistent cutaway slice of the MHC binding cleft (displayed as a grey mesh) which best indicates the binding pockets for the P1/P5/PC positions (side view - pockets A, E, F) and for the P2/P3/PC-2 positions (top view - pockets B, C, D). In some cases peptides will use a different pocket for a specific peptide position (atypical anchoring). On some structures the peptide may appear to sterically clash with a pocket. This is an artefact of picking a standardised slice of the cleft and overlaying the peptide.
Peptide neighbours
|
P1
GLY
GLU63
TRP167
MET5
TYR171
TYR159
TYR7
LYS66
TYR59
|
P2
ILE
TYR7
HIS70
TYR99
PHE9
LYS66
MET45
VAL67
GLU63
TRP167
TYR159
|
P3
LEU
TYR159
LEU156
LYS66
HIS114
HIS70
TYR99
ARG97
|
P4
GLU
LYS66
|
P5
PHE
LEU156
VAL152
GLN155
|
P6
VAL
ALA69
LYS66
HIS70
THR73
|
P7
PHE
HIS114
VAL152
ARG97
LEU156
TRP147
TYR116
THR73
ASP77
|
P8
THR
VAL76
LYS146
TRP147
THR73
ASP77
|
P9
LEU
ILE124
TRP147
THR143
TYR116
LEU81
TYR84
ASP77
TYR123
THR80
LYS146
|
Colour key
Data provenance
Neighbours are calculated by finding residues with atoms within 5Å of each other using BioPython Neighboursearch module. The list of neighbours is then sorted and filtered to inlcude only neighbours where between the peptide and the MHC Class I alpha chain.
Colours selected to match the YRB scheme. [https://www.frontiersin.org/articles/10.3389/fmolb.2015.00056/full]

|
A Pocket
TYR159
THR163
TRP167
TYR171
MET5
TYR59
GLU63
LYS66
TYR7
|
B Pocket
ALA24
VAL34
MET45
GLU63
LYS66
VAL67
TYR7
HIS70
PHE9
TYR99
|
C Pocket
HIS70
THR73
HIS74
PHE9
ARG97
|
D Pocket
HIS114
GLN155
LEU156
TYR159
LEU160
TYR99
|
E Pocket
HIS114
TRP147
VAL152
LEU156
ARG97
|
F Pocket
TYR116
TYR123
THR143
LYS146
TRP147
ASP77
THR80
LEU81
TYR84
VAL95
|
Colour key
Data provenance
|
1. Beta 2 microglobulin
Beta 2 microglobulin
|
10 20 30 40 50 60
MIQRTPKIQVYSRHPAENGKSNFLNCYVSGFHPSDIEVDLLKNGERIEKVEHSDLSFSKD 70 80 90 WSFYLLYYTEFTPTEKDEYACRVNHVTLSQPKIVKWDRDM |
|
2. Class I alpha
HLA-A*02:01
IPD-IMGT/HLA
[ipd-imgt:HLA35266] |
10 20 30 40 50 60
GSHSMRYFFTSVSRPGRGEPRFIAVGYVDDTQFVRFDSDAASQRMEPRAPWIEQEGPEYW 70 80 90 100 110 120 DGETRKVKAHSQTHRVDLGTLRGYYNQSEAGSHTVQRMYGCDVGSDWRFLRGYHQYAYDG 130 140 150 160 170 180 KDYIALKEDLRSWTAADMAAQTTKHKWEAAHVAEQLRAYLEGTCVEWLRRYLENGKETLQ 190 200 210 220 230 240 RTDAPKTHMTHHAVSDHEATLRCWALSFYPAEITLTWQRDGEDQTQDTELVETRPAGDGT 250 260 270 FQKWAAVVVPSGQEQRYTCHVQHEGLPKPLTLRWEP |
|
3. Peptide
|
GILEFVFTL
|
|
4. T cell receptor alpha
T cell receptor alpha
TRAV27
|
10 20 30 40 50 60
QLLEQSPQFLSIQEGENLTVYCNSSSVFSSLQWYRQEPGEGPVLLVTVVTGGEVKKLKRL 70 80 90 100 110 120 TFQFGDARKDSSLHITAAQPGDTGLYLCAGAGSQGNLIFGKGTKLSVKPNIQNPDPAVYQ 130 140 150 160 170 180 LRDSKSSDKSVCLFTDFDSQTNVSQSKDSDVYITDKCVLDMRSMDFKSNSAVAWSNKSDF 190 ACANAFNNSIIPEDTFFPS |
|
5. T cell receptor beta
T cell receptor beta
TRBV19
|
10 20 30 40 50 60
GGITQSPKYLFRKEGQNVTLSCEQNLNHDAMYWYRQDPGQGLRLIYYSQIVNDFQKGDIA 70 80 90 100 110 120 EGYSVSREKKESFPLTVTSAQKNPTAFYLCASSIRSSYEQYFGPGTRLTVTEDLKNVFPP 130 140 150 160 170 180 EVAVFEPSEAEISHTQKATLVCLATGFYPDHVELSWWVNGKEVHSGVCTDPQPLKEQPAL 190 200 210 220 230 240 NDSRYALSSRLRVSATFWQDPRNHFRCQVQFYGLSENDEWTQDRAKPVTQIVSAEAWGRA D |
Data provenance
Sequences are retrieved via the Uniprot method of the RSCB REST API. Sequences are then compared to those derived from the PDB file and matched against sequences retrieved from the IPD-IMGT/HLA database for human sequences, or the IPD-MHC database for other species. Mouse sequences are matched against FASTA files from Uniprot. Sequences for the mature extracellular protein (signal petide and cytoplasmic tail removed) are compared to identical length sequences from the datasources mentioned before using either exact matching or Levenshtein distance based matching.
Downloadable data
Components
Data license
Footnotes
- Protein Data Bank Europe - Coordinate Server
- 1HHK - HLA-A*02:01 binding LLFGYPVYV at 2.5Å resolution - PDB entry for 1HHK
- Protein structure alignment by incremental combinatorial extension (CE) of the optimal path. - PyMol CEALIGN Method - Publication
- PyMol - PyMol.org/pymol
- Levenshtein distance - Wikipedia entry
- Protein Data Bank Europe REST API - Molecules endpoint
- 3Dmol.js: molecular visualization with WebGL - 3DMol.js - Publication
- Protein Data Bank Europe REST API - Publication endpoint
- PubMed Central Europe REST API - Articles endpoint

This work is licensed under a Creative Commons Attribution 4.0 International License.